Marine Barcoding Synthesis workshop
Long Beach, CA
February 5, 2009
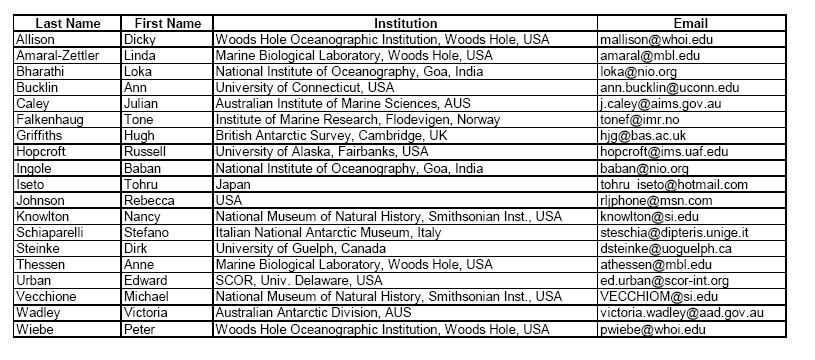
Attendees:
AGENDA
9:00 Introductions
Presentations
9:10 MarBoL Overview – Dirk Steinke
9:25 MarBoL Workshops for Spring 2009 – Ann Bucklin
CoML Project Updates
9:30 CMarZ – Ann Bucklin
9:40 CAML – Victoria Wadley
9:50 CReefs and BioCode/MOOREA – Nancy Knowlton
10:00 ICOMM – Linda Amaral-Zettler
10:10 Indian Ocean – Baban Ingole
Discussion Questions
- How many marine barcodes have been determined and by whom?
- How many marine species will be barcoded by October 2010?
- How can we accelerate and optimize our progress in marine barcoding?
- For which applications is COI best?
- CoML efforts toward a leafy tree of life: what is being done now, and what will be completed by October 2010?
- What are the most promising and realistic approaches for rapid DNA-based biodiversity surveys of marine environments?
Welcome (Ann Bucklin)
The meeting opened with a description of the goals of the meeting, including information about the workshops that MarBoL will be sponsoring. MarBoL will accelerate barcoding through coordination, with a particular focus on CoML projects. How can we help each other? Attendees introduced themselves.
Introduction to MarBoL (Dirk Steinke)
MarBoL has been ongoing for about one year now. MarBoL will accelerate marine barcoding. First question was where they started.
Main objectives are to:
1) Identify and sequence large barcodeable collections from three museums; solve problems (e.g., sponges). Need to have 50,000 barcodes by 2010.
2) Make barcoding available to CoML researchers
3) Knowledge transfer: info on new primers as soon as new routine is working.
4) The tree of Marine Life.
Update on MarBoL:
1) Start of MarBoL in January 2008 ($1M from Sloan Foundation)
2) CoML barcoding: six visitors to CCDB and SI in 2008; 2 in early 2009 to CCDB.
3) Museums started sampling, digitizing process
4) Approximately 12,000 marine species barcoded.
5) Participation at meetings and barcoding workshops.
Ongoing efforts:
1) Assembly of best practice protocols and dissemination.
1) Proposal writing: SharkBoL; CephBoL; OceanObs09; iBoL marine bio surveillance.
3) Support CoML field projects barcoding programs.
4) Identification of additional collections. (i.e. Russian collections)
5) Extending web site
6) Facilitate upload of barcoding data to public databases (BoLD, GenBank).
Discussion
Lack of verification of barcode data: identifications are not verified; no metadata for barcoded animals; geospatial data (collection location) and voucher specimen are needed. Recommend submitters use GenBank “structured comments”, which can include the metadata. Also possible to request metadata from the GenBank submitters (ICOMM and CAML have done this); very time-consuming. Also discussed consequences of bad identifications; how to determine the correct barcode when more than one has been submitted. BoLD is better in this regard, with confirmation of taxonomy and requirements for metadata. There are checkout procedures done by BoLD for newly-submitted sequences; identification engine with several layers in BoLD.
How many marine barcodes? Dirk estimates 12,000 species, but BoLD database gives number as ~4,000, since does not include GenBank sequences and does not include invertebrates. BoLD list marine fishes, birds, and mammals only. In order to judge completeness, CoML projects should send species lists to Dirk, which would be very helpful to BoLD.
CMarZ Barcoding Update (Ann Bucklin)
Three approaches: barcoding of identified specimens for target taxonomic groups, barcoding of all specimens collected during a field campaign or within a particular ocean region, and environmental sequencing of unsorted samples. Environmental sequencing (or community metagenetics) may have many applications for ocean observation, rapid biodiversity assessments, and ecosystem monitoring.
CAML Barcoding Update (Victoria Wadley)
Barcoding for CAML started in 2005. CAML now has a dedicated barcoding coordinator (Rachel Brandt).. Region has 14,000 taxa, of which about half validated. CAML studies microbes to whales. Currently have barcodes for 3,000 species and may have up to 10,000 by end of 2010. Learning how to assemble the material and get it ready for barcoding that includes the metadata; will include sequence data and geo-referenced biological data. Plan to sequence up to 8 genes for each specimen. Also take snapshots of all specimens. Demonstration projects, including octopus data analysis using GeoPhyloBuilder. Proposal to barcode all squid has been funded.
Discussion
Data discovery and coordination: Researchers cannot see data on BoLD; data discovery should be improved to allow coordination among researchers from different projects, ships, countries, etc. CoML mapping and visualization efforts will help foster communication.
CReefs Update (Nancy Knowlton)
Barcoding mandatory for papers, but tiny drop in the bucket. Her group has very large number of undetermined species; thousands of species ready for barcoding. Corals, sea anemones, and sponges don’t work for COI. Problems with coral hybridization and saying what a species is. Particular focus on environmental sequencing: resolve technical problems associated with size range of organisms; quantitative interpretation of DNA results. Will use environmental sequencing to determine biodiversity when species names unknown; determine diversity on ARMS after colonization.
Discussion
Do ARMS give accurate biodiversity assessment? ARMS (Autonomous reef monitoring structure) capture larvae that settle on clean hard substrate, but not everything. Leave out for year or more to increase species colonizing. Need to have the chambers widely distributed; will also ground-truth.
Does environmental sequencing give useful view of biodiversity?: If only know that genes have changed, what is the significance of the change? Can say what groups – if not species – are present and changing. Useful for marine monitoring and observation systems. May be used for CPR samples? Useful to see range shifts in species boundaries. Need environmental data along with the DNA data. Useful for offshore oil platforms, which have mandatory monitoring in water column and bottom; also for harbors that need monitoring. Consider environmental sequencing of COI for species, 18S for groups, etc. Concerns about counting species from COI for groups with prevalent pseudogenes (e.g., copepods); build cDNA library for COI (but concerns about preservation in RNAlater). Need effort in data interpretation for species diversity estimates; despite possible inaccuracies, still will get order of magnitude more information.
ICOMM Barcoding Update (Linda Amaral-Zettler)
ICOMM includes bacteria, archaea, eukarya, and viruses. Uses a Tag sequencing strategy (V6 region) to count different kinds of microbes in a community; useful for quantification, since V6 for bacteria has only a few copies; not useful as a phylogenetic tool. Studying microbial population structure of the world’s oceans, working with CoML field projects and LTER sites. Use V9 region for Eukarya. 454 sequencing will allow longer sequences soon.
Examples of ICOMM field projects: 1) MIRADA Deep Sequencing: pilot in Mount Hope Bay is focusing on Eucarya, with legacy data on the ICOMM web site and MICROBIS database; 2) BioMarks EU project coordinated by Colomban de Vargas; 3) POSEIDON plans 45 million 454 Tags; 4) TARA Oceans cruise funded by Eric Karsenti.
Indian Ocean Barcoing (Baban Ingole)
Indian Ocean region includes 90,000 records in OBIS. Training program in India for (commercial) fish genetics; attracted funding from other agencies for analysis of other groups. Already starting to analyze some zooplankton species, chaetognaths and euphausiids. Consistent with CoML and CBoL efforts to build capacity throughout the world.
Discussion
Data discovery issues: For zooplankton, CMarZ is establishing a CMarZ Barcode Network, so members can access lists of zooplankton species that have been collected and barcoded by CMarZ laboratories. CMarZ is also identifying a “barcoding coordinator” for each taxon, to help coordinate and accelerate barcoding efforts toward completion.
Formalin preserved material: Recommendation to review a USA National Academy of Sciences workshop report on DNA analysis from formalinized tissue. Report says that short DNA sequences are possible from tissue preserved in pH-buffered formalin. Tissue preserved in formalin and moved to alcohol is usually fine for DNA sequencing. Discussion of new projects to examine formalin preserved tissue in benthic collections using 454 sequencing technology. Possible proposal to CBoL for this study?
Victoria thanked Ann for hosting this workshop.
Adjourned at 12:00 noon.